SUMMARY
This article evaluates the presence of bacteria from the Enterobacteriaceae family (i.e. total coliforms, thermotolerant coliforms, and Escherichia coli (E. coli) in different types of reused water (grey water, rain water, and reclaimed water from urban water features) and problems with their detection (especially sensitivity and specificity of cultivation methods). The species composition of coliform bacteria in purified grey water with their antibiotic resistance are presented. As expected, the greatest antibiotic resistance was found in penicillin derivatives (ampicillin and amoxicillin clavulanate), which accounted for 32 % of the E. coli/Shigella group. Resistance to other studied antibiotics was demonstrated in 7–12 % of all coliform strains. The most resistant strains were isolated from medical facilities. Only one strain (E. coli from a medical facility) was resistant to meropenem.
INTRODUCTION
Escherichia coli (E. coli) is the most important indicator of faecal contamination and has practically replaced previously used coliform bacteria (or thermotolerant/faecal coliform bacteria) which, unlike E. coli, may not always have faecal origin and some representatives may reproduce normally in the aquatic environment. E. coli is thus a key microbiological indicator in most regulations, which include requirements for water hygienic safety. This is no different in the case of reused water. During our search of foreign regulations concerning requirements for the quality of reused water and rainwater when used indoors, we found that 12 of the 14 regulations monitor E. coli, 4 regulations also prescribe monitoring for coliform bacteria, and 3 require monitoring of faecal (thermotolerant) coliform bacteria either as the sole microbiological marker or together with E. coli. Matějů et al. [1] compiled an overview of 22 standards and legal regulations to regulate water reuse, including irrigation water. Each of these regulations sets out requirements for microbiological indicators, represented by total coliforms, thermotolerant coliforms, and E. coli. Interestingly, E. coli (59 %) is most often mentioned (i.e., 13 times), followed by coliform bacteria and faecal (thermotolerant) coliforms (seven times in total). Of these, a combination of E. coli and coliform bacteria, and once a combination of coliform and faecal (thermotolerant) coliform bacteria, are mentioned four times. Occasionally, the term “thermotolerant coli” appears in a text, discussion, or draft regulation, which is misleading and probably refers to a group of thermotolerant coliforms. As such, E. coli is thermotolerant, but this property is the first to be lost in an aqueous (non-physiological) environment.
The subject of this article is not to evaluate the significance of these indicators in reused water, but to show their heterogeneity in this environment and methodological difficulties in their determination. These are mainly due to the complex, highly active, and heterogeneous matrix (high content of the accompanying microflora). In order for the results obtained to be relevant (i.e., correct and accurate), the methods used must be sufficiently sensitive and selective, which is often a big problem for such active matrices. We have already addressed this issue in the case of natural bathing water [2], where the most suitable method for the determination of E. coli proved to be the financially demanding MPN (most probable number) procedure according to ČSN EN ISO 9308-2.
METHODOLOGY
Samples and their collection
From June 2020 to December 2021, in compliance with the applicable rules for sampling (ČSN EN ISO 5667-1, ČSN EN ISO 5667-3, ČSN EN ISO 19458), samples of grey water (residential buildings, public buildings, and medical facilities), rainwater (residential buildings and public buildings) and reused water from urban water features were collected in sterile sample boxes. The samples were transported under constant cooling (5 ± 3 °C) and processed within a maximum of 18 hours after collection.
Methods of analyses
Methods for culture determination were used according to the following standards and procedures:
- ČSN EN ISO 9308-1 – Chromocult Coliform Agar (Merck), membrane filtration (filters with a porosity of 0.45 µm), cultivation for 24 hours at 36 °C.
- ČSN EN ISO 9308-2 (Colilert/Quanti-Tray),
cultivation for 18 hours or 24 hours (depending on the type of test) at 36 °C. - ČSN 75 7835 – m-FC Agar (Merck), membrane filtration
(filters with a porosity of 0.45 µm), cultivation for 24 hours at 44 °C. - Optimized method for E. coli determination on chromogenic TBX medium (Merck), membrane filtration (filters with a porosity of 0.45 µm), cultivation for 24 hours at temperatures of 36 °C or 44 °C, possible pre-cultivation on selective medium TSA/CA or TBX for 4 hours at 36 °C.
Determination of antibiotic resistance
145 strains of gram-negative bacteria from the family Enterobacteriaceae, belonging to the group of coliform bacteria, were isolated from purified grey water (11 localities). The strains were purified and identified by MALDI-TOF and tested for antibiotic susceptibility by disk diffusion according to the European Committee on Antimicrobial Susceptibility Testing, version 7.0 of January 2019, so-called EUCAST [3] (Fig. 1). If a low number of colonies (less than five) were detected in the sample, all were tested; otherwise, five to eight colonies were selected, preferably of different appearance. The strains were tested for antibiotics from the Oxoid company (disc content in parentheses): ampicillin (10 µg); amoxicillin-clavulanic acid (30 µg); cefotaxime (5 µg); cefepime (30 µg); ceftazidime (10 µg); meropenem (10 µg); ciprofloxacin (5 µg) and gentamicin (10 µg).
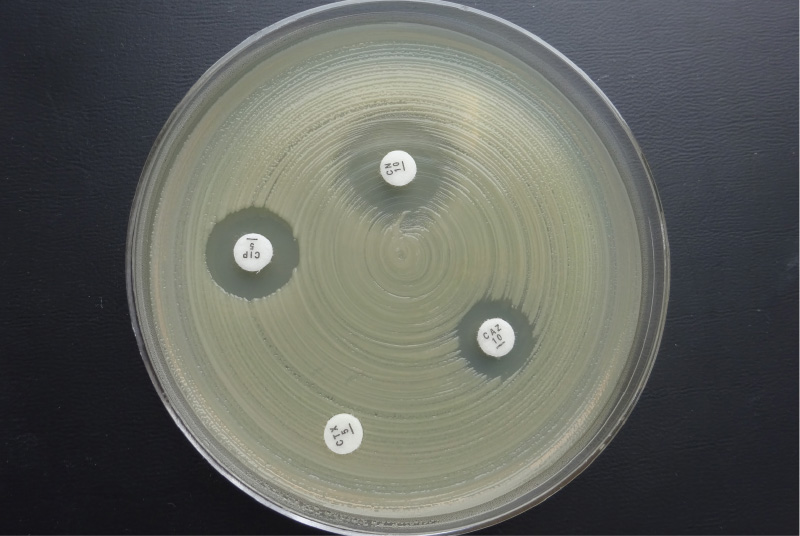
Fig. 1. Disc diffusion method – disks impregnated with the antibiotics are placed in a Petri dish with Mueller Hinton medium inoculated with the test strain; based on the size of the growth inhibition zone around the disc, susceptibility/resistance to a given antibiotic is determined.
RESULTS AND DISCUSSION
Representation of E. coli among coliform bacteria in various types of water (not only reused)
Like other coliform bacteria, E. coli is a member of the Enterobacteriaceae family; however, unlike other species, it does not survive in the aquatic environment for long periods of time. The percentage of E. coli among coliform (or thermotolerant coliform) bacteria may indicate the severity of direct faecal contamination and, in part, the level of organic contamination. Tab. 1 shows the different percentages of E. coli among coliform bacteria in the different types of reused water we monitored. Only the results of sampling in 2021 are given, as all these analyses were performed by the method according to ČSN EN ISO 9308-2, and are therefore comparable. For statistical purposes, values above the working range > 2 419.2 were replaced by 3 750 and values > 4 838.4 by 7 500 MPN/100 ml.
Tab. 1. E. coli among total coliform bacteria in different types of reused water
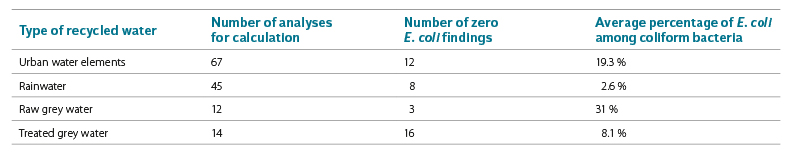
As expected, the most identified representatives of E. coli among coliform bacteria were found in raw grey water (31 %). However, these results are also the least accurate, as most results were above the working range
of the method. Significantly less E. coli was found in treated grey water (8.1 %).
The second highest incidence of E. coli was found in urban water elements, which is logical because direct faecal pollution can also occur there. In contrast, the least E. coli among coliform bacteria (2.6 %) was determined in rain water, where the absolute numbers were also low. For comparison, e.g. in surface water (the Labe and Vltava rivers), E. coli was found among coliform bacteria in 24.7 % of cases [4]. Here, however, the indicators were not determined by the method according to ČSN EN ISO 9308-2, but according to ČSN 75 7837 (Endo agar), or ČSN 75 7835 (m-FC with β-D-glucuronidase). A change in the method for determining coliform bacteria, in this case fermentation of lactose versus β-D-galactosidase activity, has led to a broader understanding of the group of coliform bacteria, which currently includes significantly more species than in the past.
Methods tested for determining coliform bacteria and E. coli in reused water
The problem of detection – especially of the species E. coli in water with rich accompanying microflora – is the fact that determination is usually performed in parallel with detection of coliform bacteria on a very sensitive medium. Detectors (membrane filters or Petri dishes) tend to be overgrown not only with the accompanying microflora, but also with coliform bacteria, which discriminates against E. coli (usually in a significant minority), so that its colonies cannot grow properly on the medium. Dilution of the samples will not help in this case because E. coli is diluted first (Fig. 2), which makes it possible to obtain a false negative result, or the limit of detection is significantly increased. Statistical evaluation of these types of results was not performed in our survey because some methods were quickly excluded after analyses of several series of samples as methods unsuitable for a given matrix.
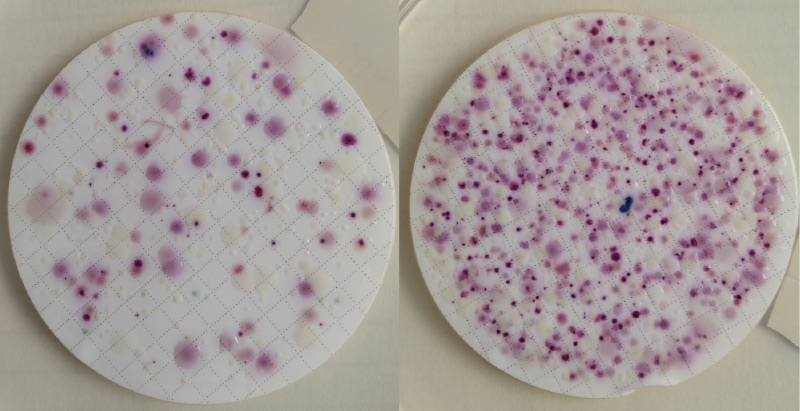
Fig. 2. Detection of E. coli according to EN ISO 9308-1 (CCA medium) from grey water matrix – presence of E. coli (blue and violet blue colonies) in higher volume is apparent, but difficult to calculate in the growth of coliform bacteria (rose/pink colonies); at higher dilutions it can no longer be detected
Another possibility is to use m-FC medium (ČSN 75 7835) and determine either thermotolerant coliform bacteria (indicator as such) or identify E. coli by subsequent detection of the enzyme β-D-glucuronidase. The first option would probably be more appropriate as m-FC medium is extremely selective (with low productivity below 50 %) and the cultivation temperature of 44 °C is very high; however, the results of E. coli determinations are often underestimated by this method [2]. Although, in most cases, the accompanying microflora is eliminated, in extremely microbially loaded samples (which contain a large number of proteolytic bacteria) it can increase and, in this case, sometimes interfere with acid-base reactions, on which this medium is based. The result is the formation of red rather than blue colonies (proteolytic bacteria can produce large amounts of ammonia, and therefore this alkaline environment may not allow the expression of acid reaction products formed after lactose fermentation).
The MPN method according to ČSN EN ISO 9308-2, which is sufficiently sensitive and selective and has a working range of three orders of magnitude, proved to be the most suitable standardized method so far. We also use this method by default in our analyses. However, it also has certain disadvantages, which is primarily its cost, and then the necessity to purchase a relatively expensive Quanti-Tray system (which is why many laboratories do not have it). Last but not least, this method produces an enormous amount of infected plastic waste. Therefore, we looked for another way to determine E. coli in reused water.
In 2019, some Croatian authors [5] published a proposal for an optimized method for the determination of E. coli in bathing water on TBX medium (Trypton Bile X-Glucuronide). TBX is a chromogenic medium where E. coli is detected by β-D-glucuronidase enzyme activity and grows in the form of turquoise colonies (all other bacteria, including other species of the Enterobacteriaceae family, grow as colourless colonies). For the determination of E. coli from natural water, pre-cultivation of inoculated dishes or membrane filters is used, lasting 4 hours at 36 °C, either on nutrient-rich non-selective medium or directly on TBX medium. The mentioned authors [5] used MMGA (Minerals Modified Glutamate Agar) for pre-cultivation, while we used tryptone soy agar (TSA) or Columbia agar (CA) as well as TBX. Our preliminary results have shown that the optimized method is promising [6] (Fig. 3). The accompanying microflora was sufficiently suppressed and the number of E. coli determined by cultivating on solid medium was comparable with the results obtained by MPN method when cultivating in liquid medium according to ČSN EN ISO 9308-2 (Quanti-Tray), which is the same principle of determination (β-D-glucuronidase).
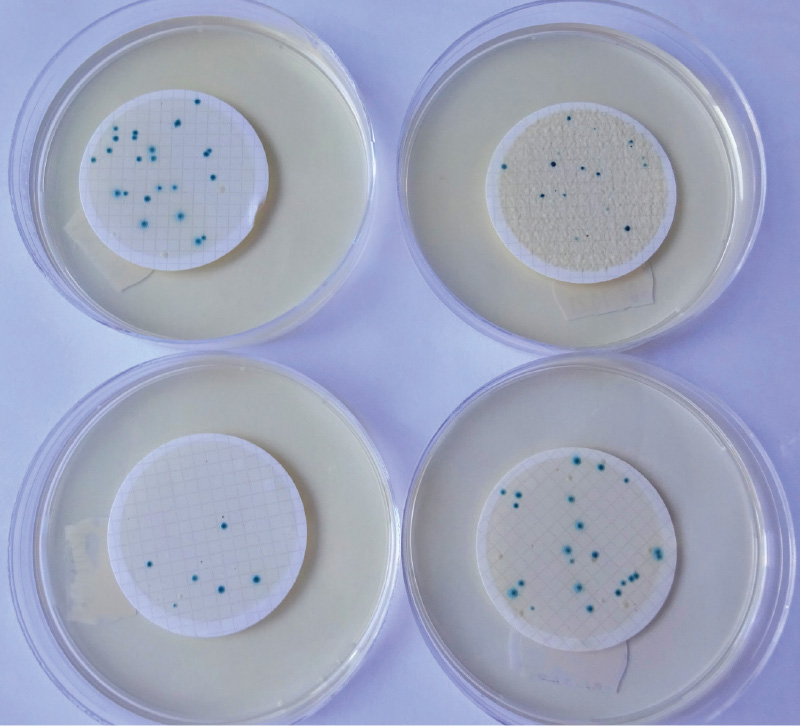
Fig. 3. E. coli on TBX medium (top left: 4 hrs cultivating at 36 °C, then for 20 hrs at 44 °C; top right: cultivating for 24 hrs at 36 °C; bottom left: incubation for 24 hrs at 44 °C; bottom right: cultivation on CA for 4 hrs at 36 °C, then for 20 hrs at 44 °C on TBX)
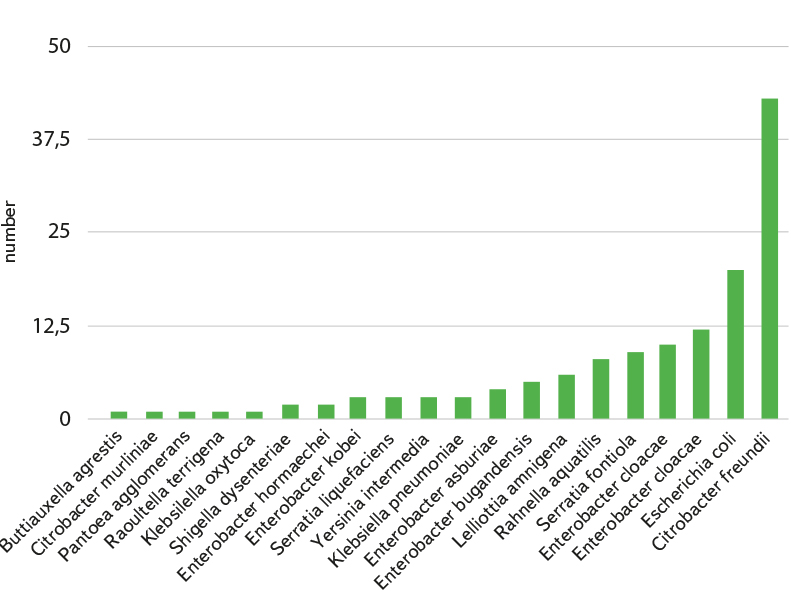
Fig. 4. List of isolated and identified coliform bacteria species
Coliform bacteria isolated from purified grey water and their antibiotic resistance
As already mentioned, 145 strains of bacteria from the family Enterobacteriaceae, belonging to the group of coliform bacteria, were isolated and identified from purified grey water (11 localities). These strains were further tested for possible antibiotic resistance. The strains achieved an average score of 2.29 when identified on MALDI-TOF, which means highly probable identification of the species. The species composition of isolated strains is shown in Fig 4. However, some species, although thus “accurately” identified, belong to broader taxonomic groups (E. coli/Shigella, Enterobacter cloacae complex, Klebsiella oxytoca/Raoultella terrigena, etc.), and their distinction is therefore rather theoretical. The assessment of antibiotic resistance is thus presented at the level of these groups or genera.
Based on data from the literature search and from the EUCAST clinical breakpoint table [7], a set of antibiotic disks suitable for resistance testing in coliform bacteria (bacteria from the Enterobacteriaceae family) was selected. The final set includes penicillins (ampicillin, amoxicillin-clavulanic acid), cephalosporins (cefotaxime, cefepime and ceftazidime), carbapenems (meropenem), fluoroquinolones (ciprofloxacin), and aminoglycosides (gentamicin). Tetracycline has not been tested because EUCAST does not provide interpretation criteria for this group of bacteria, so-called clinical breakpoints, which are the averages of zones according to which strains are categorized as sensitive or resistant.
As expected, the highest resistance was found for penicillin derivatives (ampicillin and amoxicillin-clavulanic acid). However, some genera of bacteria are naturally resistant to these penicillins [8]. Of the strains we isolated, in the case of ampicillin, it was mainly the genera Citrobacter, Enterobacter, Klebsiella, and Raoultella; in the case of amoxicillin-clavulanic acid it was Citrobacter freundii and the genus Enterobacter. From the point of view of our research, E. coli, which does not have a natural resistance to these antibiotics, is especially important. Of the 22 E. coli strains tested (along with Shigella dysenteriae), 32 % were resistant to both ampicillin and amoxicillin-clavulanic acid. Our results are relatively consistent with the literature data, according to which ampicillin resistance was found in 38 % of E. coli strains from wastewater [9] and in 18 % of E. coli strains isolated from wastewater and sludge [10].
Meropenem resistance was demonstrated in only one E. coli strain from a medical facility (i.e., 0.7 %), while 11.7 % of all coliform strains were resistant to ciprofloxacin, 9 % to gentamicin, 7 % to cefepime, 9 % to ceftazidime, and 9 % to cefotaxime. For comparison with other studies worldwide [9], 6 % of ciprofloxacin-resistant E. coli isolates from wastewater were found. Strains of bacteria from the family Enterobacteriaceae (especially the genera Enterobacter and Citrobacter) resistant to gentamicin, ciprofloxacin, meropenem, ceftazidime, and amoxicillin-clavulanic acid have also been isolated from well water in Slovakia [11].
Demonstrated resistances in individual genera of bacteria or monitored types of buildings with reused water are shown in Fig. 5 and 6. The highest level of antibiotic resistance was found in strains isolated from a medical facility. Unfortunately, we have the smallest number of these strains. Fourteen strains (9.7 %) were resistant to three or more tested antibiotics, so they were multi-resistant strains.
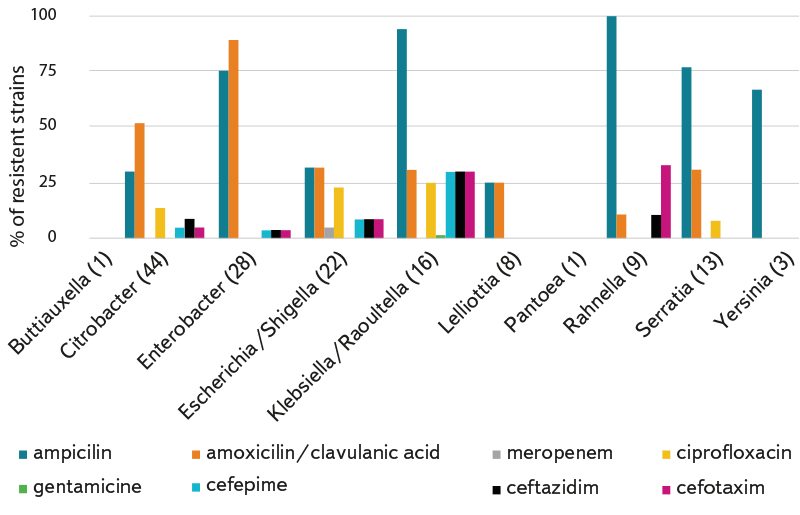
Fig. 5. Percentage of resistant strains in individual genera/taxonomic groups of coliform bacteria; the number of isolated and tested strains of the relevant genus/group is given in brackets after the genus name.
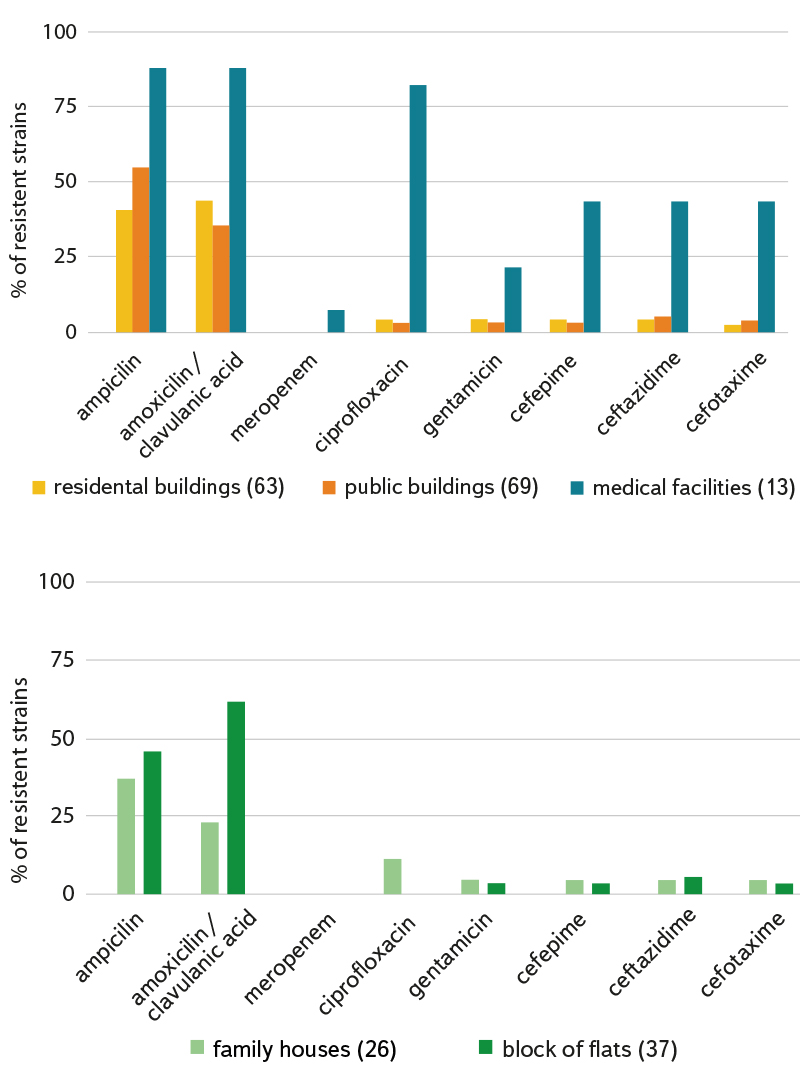
Fig. 6a (top) and 6b (bottom). Percentage of resistant strains (without genus distinction) according to isolation from different types of localities; the number of isolated and tested strains is given in brackets after the type of locality; we consider family houses to be buildings with a maximum of two flats.
CONCLUSION
Among coliform bacteria, E. coli occurs in reused water and rainwater from 2.6 % (rainwater) to 31 % (raw grey water). Due to the high content
of the accompanying microflora (especially in grey water), the method according to ČSN EN ISO 9308-2 is currently the most suitable method for E. coli detection. For the future, there seems to be a promising optimized method for the determination of (only) E. coli (with a four-hour resuscitation step) on TBX medium.
Using the MALDI-TOF method, 19 species of coliform bacteria occurring in purified grey water were identified and tested for antibiotic susceptibility. In terms of antibiotic resistance, the highest was, as expected, identified for penicillin derivatives (ampicillin and amoxicillin-clavulanic acid), at 32 %
in the E. coli/Shigella group. Resistance to other monitored antibiotics was demonstrated in 7–12 % of all strains of coliform bacteria. The most resistant strains were isolated from medical facilities. Only one E. coli strain isolated from water from a medical facility was resistant to meropenem. These results are consistent with the results of other experts who have isolated bacteria from different types of water (wastewater, sludge, wells) [9–11].
Acknowledgements
This article was written within the project TA CR SS01010179 “Determination of hygienic requirements for reused water used in buildings and urban water features”.
The article has been peer-reviewed.
